Energy Volume Curve Agent#
Based on the previous tutorial, the next step is to extend the simple crystal structure agent to address the benchmark challenge of calculating an energy-versus-volume curve using the EMT simulation code. Again, this demonstration is based on the Langchain tutorial for custom agents.
As a first step, we import a number of python modules, these consist of the ASE, some standard python libraries as well as a number of tools from langchain:
from ase.atoms import Atoms
from ase.build import bulk
from ase.calculators.emt import EMT
from ase.constraints import UnitCellFilter
from ase.eos import calculate_eos, plot
from ase.optimize import LBFGS
from getpass import getpass
from typing import List
from langchain_core.tools import tool
from langchain.agents import AgentExecutor
from langchain.agents.format_scratchpad.openai_tools import format_to_openai_tool_messages
from langchain.agents.output_parsers.openai_tools import OpenAIToolsAgentOutputParser
from langchain_core.pydantic_v1 import BaseModel
from langchain_core.prompts import ChatPromptTemplate, MessagesPlaceholder
from langchain_openai import ChatOpenAI
Multiple Functions#
To connect multiple python functions, it is essential to convert all inputs and outputs to JSON compatible data types as the communication to the LLM in the background happens in terms of web requests using the JSON format. This especially applies to python functions which return python objects. In the case of ASE a typical python object is the ase.atoms.Atoms() object. To convert the Atoms() object to JSON format, we use a pydantic data class as suggested by langchain:
class AtomsDict(BaseModel):
numbers: List[int]
positions: List[List[float]]
cell: List[List[float]]
pbc: List[bool]
AtomsDict.schema()
{'title': 'AtomsDict',
'type': 'object',
'properties': {'numbers': {'title': 'Numbers',
'type': 'array',
'items': {'type': 'integer'}},
'positions': {'title': 'Positions',
'type': 'array',
'items': {'type': 'array', 'items': {'type': 'number'}}},
'cell': {'title': 'Cell',
'type': 'array',
'items': {'type': 'array', 'items': {'type': 'number'}}},
'pbc': {'title': 'Pbc', 'type': 'array', 'items': {'type': 'boolean'}}},
'required': ['numbers', 'positions', 'cell', 'pbc']}
In terms of functions, two functions are defined: A get_equilibirum_lattice() function which for a given chemical element returns the optimized equilibrium structure as AtomsDict. The second function is a plot_equation_of_state() function, which takes the already optimized structure as AtomsDict as input and then uses the ASE internal functionality to plot the energy volume curve.
The important point here is that get_equilibirum_lattice() would typically just return an ase.atoms.Atoms() object, but for compatibility with the LLM it has to be converted to an AtomsDict which can be converted to JSON. Finally, in the plot_equation_of_state() function the AtomsDict is again converted to an ase.atoms.Atoms() object to continue the calculation. This is currently a bit tedious.
@tool
def get_equilibirum_lattice(chemical_symbol: str) -> AtomsDict:
"""Returns equilibrium atoms dictionary for a given chemical symbol"""
atoms = bulk(name=chemical_symbol)
atoms.calc = EMT()
ase_optimizer_obj = LBFGS(UnitCellFilter(atoms))
ase_optimizer_obj.run(fmax=0.000001)
return AtomsDict(**{k: v.tolist() for k, v in atoms.todict().items()})
@tool
def plot_equation_of_state(atom_dict: AtomsDict) -> str:
"""Returns plot of equation of state of chemical symbol for a given atoms dictionary"""
atoms = Atoms(**atom_dict.dict())
atoms.calc = EMT()
eos = calculate_eos(atoms)
plotdata = eos.getplotdata()
return plot(*plotdata)
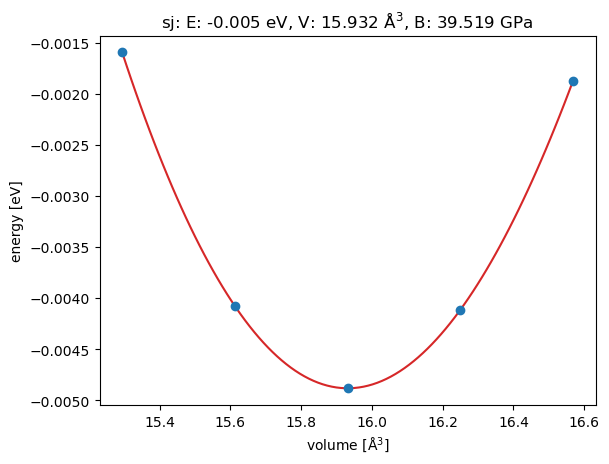
Finally, the functions converted to LLM tools using the tool decorator can again be tested using the invoke() function. It takes a python dictionary as input to address the different arguments individually. With this test, the correct implementation of the python functions is validated:
el = "Al"
structure_dict = get_equilibirum_lattice.invoke({"chemical_symbol": el})
plot_equation_of_state.invoke({"atom_dict": structure_dict})
Step Time Energy fmax
LBFGS: 0 18:48:16 -0.001502 0.161794
LBFGS: 1 18:48:16 -0.002533 0.135471
LBFGS: 2 18:48:16 -0.004879 0.005452
LBFGS: 3 18:48:16 -0.004883 0.000157
LBFGS: 4 18:48:16 -0.004883 0.000000
/var/folders/z7/3vhrmssx60v240x_ndq448h80000gn/T/ipykernel_55498/2598102435.py:6: FutureWarning: Import UnitCellFilter from ase.filters
ase_optimizer_obj = LBFGS(UnitCellFilter(atoms))
<Axes: title={'center': 'sj: E: -0.005 eV, V: 15.932 Å$^3$, B: 39.519 GPa'}, xlabel='volume [Å$^3$]', ylabel='energy [eV]'>
Agents#
Following the same Langchain tutorial for custom agents as before a LLM agent is constructed which combines the system prompt, the python functions get_equilibirum_lattice() and plot_equation_of_state() as tools and the OpenAIToolsAgentOutputParser():
OPENAI_API_KEY = getpass(prompt='Enter your OpenAI Token:')
llm = ChatOpenAI(model="gpt-3.5-turbo", temperature=0, openai_api_key=OPENAI_API_KEY)
tools = [get_equilibirum_lattice, plot_equation_of_state]
prompt = ChatPromptTemplate.from_messages(
[
(
"system",
"You are very powerful assistant, but don't know current events. For each query vailidate that it contains a chemical element and otherwise cancel.",
),
("user", "{input}"),
MessagesPlaceholder(variable_name="agent_scratchpad"),
]
)
agent = (
{
"input": lambda x: x["input"],
"agent_scratchpad": lambda x: format_to_openai_tool_messages(
x["intermediate_steps"]
),
}
| prompt
| llm.bind_tools(tools)
| OpenAIToolsAgentOutputParser()
)
agent_executor = AgentExecutor(agent=agent, tools=tools, verbose=True)
User Interactions#
The user can then interact with the agent by asking it to Plot the equation of state of gold. In the background the agent first calls the get_equilibirum_lattice() with the parameter Au for the chemical symbol of gold and afterwards plot_equation_of_state() with the atomistic structure returned by get_equilibirum_lattice().
lst = list(agent_executor.stream({"input": "Plot the equation of state of gold"}))
> Entering new AgentExecutor chain...
Invoking: `get_equilibirum_lattice` with `{'chemical_symbol': 'Au'}`
Step Time Energy fmax
LBFGS: 0 18:48:23 0.002606 0.308859
LBFGS: 1 18:48:23 0.000032 0.077808
LBFGS: 2 18:48:24 -0.000135 0.003099
LBFGS: 3 18:48:24 -0.000135 0.000029
/var/folders/z7/3vhrmssx60v240x_ndq448h80000gn/T/ipykernel_55498/2598102435.py:6: FutureWarning: Import UnitCellFilter from ase.filters
ase_optimizer_obj = LBFGS(UnitCellFilter(atoms))
LBFGS: 4 18:48:24 -0.000135 0.000000
numbers=[79] positions=[[4.761270571021482e-17, -3.4431732128676475e-17, -2.0729599738876008e-16]] cell=[[7.040904860568554e-17, 2.028082809705617, 2.0280828097056176], [2.028082809705617, 1.0384771021574885e-16, 2.0280828097056176], [2.028082809705617, 2.028082809705618, 4.963320342464553e-17]] pbc=[True, True, True]
Invoking: `plot_equation_of_state` with `{'atom_dict': {'numbers': [79], 'positions': [[4.761270571021482e-17, -3.4431732128676475e-17, -2.0729599738876008e-16]], 'cell': [[7.040904860568554e-17, 2.028082809705617, 2.0280828097056176], [2.028082809705617, 1.0384771021574885e-16, 2.0280828097056176], [2.028082809705617, 2.028082809705618, 4.963320342464553e-17]], 'pbc': [True, True, True]}}`
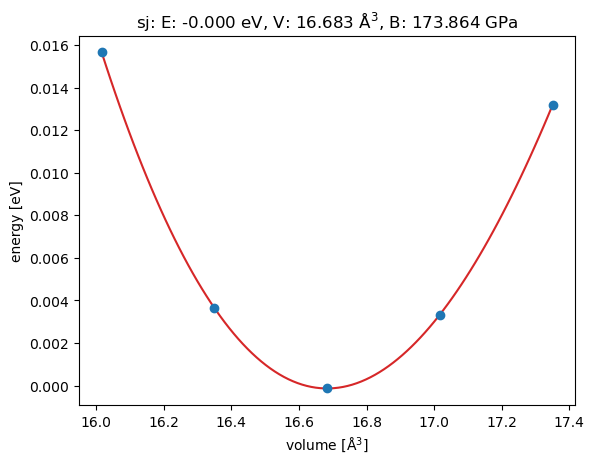
Axes(0.125,0.11;0.775x0.77)Here is the plot of the equation of state of gold.
> Finished chain.
Summary#
The important point in this example is, that the order of execution of the Python functions is not defined by the user. Instead the LLM automatically determines that plot_equation_of_state() needs an AtomsDict() object as input and that get_equilibirum_lattice() returns such a AtomsDict() object as an output, so it makes sense to call get_equilibirum_lattice() first and plot_equation_of_state() second. The same principles apply to LLM agents with a larger number of python functions.
The limiting point at the moment is that the LLMs are web services, so all Python objects have to be converted to JSON to be communicated to the LLMs. This restricts the choice of Python objects and requires the development of specialized data classes to construct those interfaces between different Python functions.
